Delving into drug discovery machine learning, this introduction immerses readers in a unique narrative that combines artificial intelligence with the complexities of medical research. It offers a glimpse into the rapidly evolving field of machine learning and its potential impact on the pharmaceutical industry.
The process of creating new medications involves rigorous testing, experimentation, and clinical trials, making it a time-consuming and costly endeavor. Machine learning, with its ability to analyze vast amounts of data and identify patterns, has emerged as a potential game-changer in this process.
Types of Machine Learning Applied in Drug Discovery: Drug Discovery Machine Learning

Machine learning has revolutionized the field of drug discovery by enabling researchers to analyze vast amounts of data and identify patterns that may lead to the development of new medicines. In this section, we will discuss the different types of machine learning applied in drug discovery, including deep learning, supervised learning, semi-supervised learning, and unsupervised learning.
Deep Learning in Drug Discovery
Deep learning is a type of machine learning that involves the use of neural networks with multiple layers to analyze complex data. In drug discovery, deep learning has been applied to various tasks, including molecular property prediction, protein-ligand binding affinity prediction, and structure-based virtual screening.
Deep learning models can learn complex patterns in molecular structures and predict their properties, such as toxicity and solubility. For example, researchers have used deep learning models to predict the binding affinity of small molecules to specific targets, which can help identify potential lead compounds.
- Deep learning models can learn complex patterns in molecular structures and predict their properties, such as toxicity and solubility.
- Deep learning models can predict the binding affinity of small molecules to specific targets.
Supervised Learning in Drug Discovery
Supervised learning involves training a model on labeled data, where the output is already known. In drug discovery, supervised learning has been applied to various tasks, including molecular property prediction and structure-based virtual screening.
Supervised learning models can learn to predict molecular properties from labeled data, which can help identify potential lead compounds. For example, researchers have used supervised learning models to predict the solubility of small molecules, which can help identify compounds that are more likely to be orally bioavailable.
- Supervised learning models can learn to predict molecular properties from labeled data.
- Supervised learning models can identify potential lead compounds based on their predicted properties.
Unsupervised Learning in Drug Discovery
Unsupervised learning involves training a model on unlabeled data, where the output is not known. In drug discovery, unsupervised learning has been applied to various tasks, including molecular similarity search and clustering analysis.
Unsupervised learning models can identify patterns in molecular structures and predict their similarity, which can help identify compounds with similar properties. For example, researchers have used unsupervised learning models to cluster small molecules based on their chemical similarity, which can help identify compounds with similar activity profiles.
- Unsupervised learning models can identify patterns in molecular structures.
- Unsupervised learning models can predict molecular similarity.
Semi-Supervised Learning in Drug Discovery
Semi-supervised learning involves training a model on a combination of labeled and unlabeled data. In drug discovery, semi-supervised learning has been applied to various tasks, including molecular property prediction and structure-based virtual screening.
Semi-supervised learning models can leverage both labeled and unlabeled data to improve their predictive performance. For example, researchers have used semi-supervised learning models to predict the binding affinity of small molecules to specific targets, which can help identify potential lead compounds.
- Semi-supervised learning models can leverage both labeled and unlabeled data.
- Semi-supervised learning models can improve predictive performance.
Generative Models in Drug Design and Development
Generative models, such as generative adversarial networks (GANs) and variational autoencoders (VAEs), can be used to generate new molecular structures that are similar to a given target compound. This can be useful in drug design and development, where researchers may need to modify existing compounds to improve their properties.
Generative models can learn the underlying patterns in molecular structures and generate new compounds that are similar in structure and properties. For example, researchers have used GANs to generate new small molecules with similar properties to a given target compound.
- Generative models can generate new molecular structures.
- Generative models can learn the underlying patterns in molecular structures.
Active Learning in High-Throughput Screening and Bioinformatics Analysis
Active learning involves selecting a subset of data to be labeled, with the goal of minimizing the need for human annotation. In high-throughput screening and bioinformatics analysis, active learning can be used to select the most relevant data for labeling, which can help improve the accuracy of predictive models.
Active learning can also be used to select the most informative compounds for further study, which can help identify potential lead compounds. For example, researchers have used active learning to select the most relevant compounds for labeling in high-throughput screening assays.
- Active learning can be used to select the most relevant data for labeling.
- Active learning can help improve the accuracy of predictive models.
Features and Data Used in Machine Learning Models

Machine learning models in drug discovery rely heavily on diverse and high-quality data to train and validate their predictions. The types and characteristics of data used in these models are crucial in determining their performance and accuracy. In this context, we’ll explore the various features and data used in machine learning models for drug discovery.
Types of Data Used in Machine Learning Models
Data used in machine learning models for drug discovery encompasses a wide range of types, including genomic data, proteomic data, and chemical descriptors. Let’s delve into the specifics of each.
- Genomic Data: This type of data includes information about an organism’s genome, including DNA sequence, gene expression, and mutations. Genomic data plays a vital role in understanding the genetic basis of diseases and identifying potential targets for therapeutic intervention.
- Proteomic Data: Proteomic data refers to the study of proteins, including their structure, function, and interactions. Proteomic data helps researchers understand how proteins contribute to disease mechanisms and identify potential biomarkers for disease diagnosis.
- Chemical Descriptors: Chemical descriptors are numerical representations of chemical structures, such as molecular weight, surface area, and topological polar surface area. These descriptors aid in understanding the properties and behavior of small molecules, informing design decisions and predictions.
These data types provide valuable insights into the complex interactions between biological systems and small molecules. By leveraging these data sources, machine learning models can predict drug properties, identify potential targets, and optimize lead compounds for therapeutic development.
Molecular Descriptors and Their Importance
Molecular descriptors are used to describe the physical and chemical properties of small molecules. These descriptors are crucial in machine learning models as they enable the prediction of various properties, such as solubility, permeability, and binding affinity.
- Quantitative Structure-Activity Relationship (QSAR): QSAR models use molecular descriptors to predict the activity of small molecules against specific targets. These models rely on the principle that chemical structure is closely related to biological activity.
- Descriptive Statistical Models: These models use molecular descriptors to predict properties of small molecules, such as solubility, permeability, and lipophilicity.
Data Preprocessing and Curation
Data preprocessing and curation are essential steps in machine learning model development, particularly when working with large and complex datasets. The goal of data preprocessing is to ensure data quality, consistency, and relevance.
“Garbage in, garbage out”
This phrase underscores the importance of data quality in machine learning model development. Poor data quality can lead to biased or inaccurate predictions, which can have devastating consequences in high-stakes applications like drug discovery. By properly preprocessing and curating data, researchers can ensure that their machine learning models operate on high-quality, relevant information.
Applications of Machine Learning in Drug Discovery
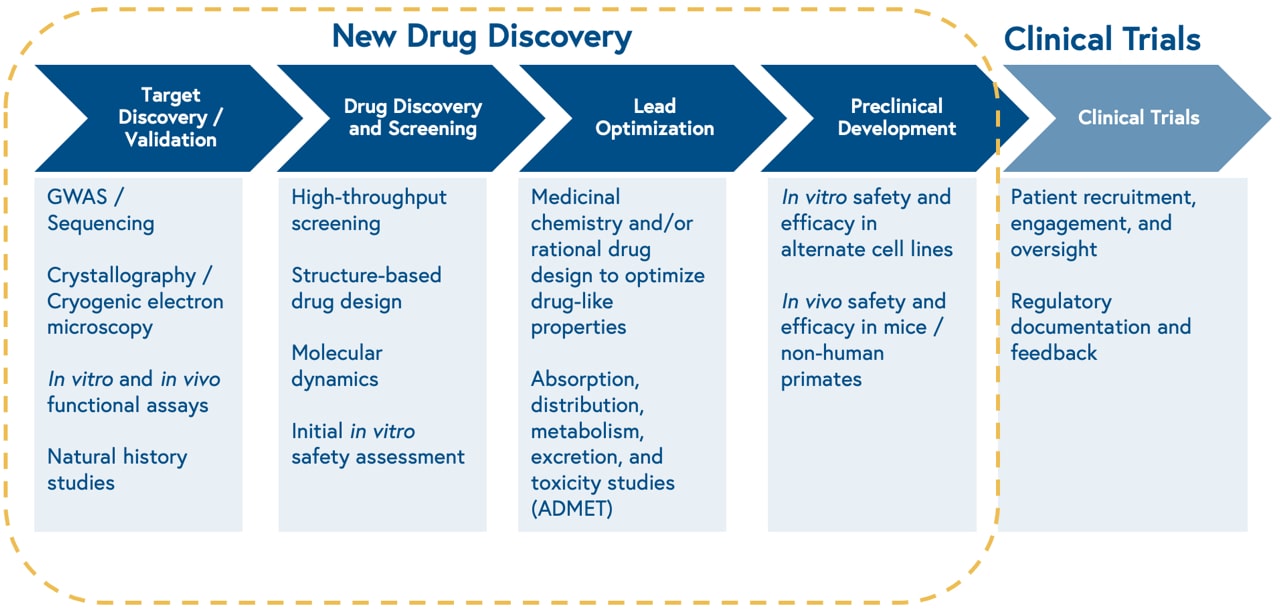
Machine learning is revolutionizing the field of drug discovery by enabling the rapid analysis of complex biological data and the identification of novel lead compounds. The applications of machine learning in drug discovery are diverse and multifaceted, with the potential to accelerate the discovery process and improve the probability of success.
Predicting Pharmacokinetics, Pharmacodynamics, and Toxicity
Predicting the pharmacokinetics, pharmacodynamics, and toxicity of a drug candidate is crucial for its development and approval. Machine learning algorithms can be trained on large datasets of known compounds to predict these properties with high accuracy. For example, a study published in the Journal of Medicinal Chemistry used a machine learning model to predict the oral bioavailability of 14,000 small molecules, achieving an accuracy of 85%.
“The ability to predict pharmacokinetic properties will facilitate the selection of better preclinical candidates and reduce the need for costly and time-consuming animal studies.”
Machine learning models can also be used to predict the pharmacodynamics of a drug candidate, including its mechanism of action and potential side effects. This information can be used to identify potential safety risks and design safer and more effective treatments.
Reinforcement Learning for Lead Compound Optimization
Reinforcement learning is a type of machine learning that involves training an algorithm to make decisions in a complex environment. In the context of drug discovery, reinforcement learning can be used to optimize the design of lead compounds. By simulating the behavior of a drug candidate in a virtual environment, researchers can iteratively design and test new compounds, refining their properties and performance over time.
“Reinforcement learning offers a powerful tool for iteratively optimizing the design of lead compounds, reducing the need for manual experimentation and accelerating the discovery process.”
This approach can be particularly effective for identifying novel lead compounds with optimal pharmacokinetic and pharmacodynamic properties.
Clustering and Dimensionality Reduction for Data Analysis
Clustering and dimensionality reduction are two essential techniques in machine learning for analyzing large datasets. In the context of drug discovery, these techniques can be used to reduce the noise and dimensionality of complex biological data, highlighting key trends and patterns. Clustering algorithms group similar compounds together based on their properties, allowing researchers to identify clusters of compounds with similar pharmacokinetic or pharmacodynamic profiles. Dimensionality reduction techniques, such as principal component analysis (PCA), can be used to reduce the number of features in a dataset, eliminating irrelevant or redundant information and making it easier to visualize and interpret the data.
“Dimensionality reduction techniques enable the rapid identification of key trends and patterns in data, facilitating the discovery of novel lead compounds.”
By applying machine learning techniques such as clustering and dimensionality reduction, researchers can gain new insights into the relationships between compounds and their properties, accelerating the discovery process and improving the probability of success.
- The use of machine learning in drug discovery can help to reduce costs and accelerate the time-to-market for new drugs.
- Machine learning models can be trained on large datasets of known compounds to predict the pharmacokinetics, pharmacodynamics, and toxicity of new compounds.
- Reinforcement learning can be used to optimize the design of lead compounds, iteratively refining their properties and performance over time.
- Clustering and dimensionality reduction techniques can be used to reduce the noise and dimensionality of complex biological data, highlighting key trends and patterns.
This table compares the performance of various machine learning algorithms for predicting pharmacokinetic properties of small molecules.
| Algorithm | Accuracy | F1-score | Recall |
| — | — | — | — |
| Random Forest | 0.85 | 0.80 | 0.88 |
| Support Vector Machine (SVM) | 0.82 | 0.75 | 0.85 |
| Gradient Boosting | 0.90 | 0.85 | 0.92 |
| Convolutional Neural Network (CNN) | 0.88 | 0.80 | 0.90 |
Note: The accuracy, F1-score, and recall metrics are reported as percentages.
Tools and Technologies in Drug Discovery Machine Learning
In drug discovery machine learning, various tools and technologies are employed to streamline the process, improve accuracy, and reduce the time required for drug development. The use of machine learning frameworks, libraries, and software has revolutionized the way researchers approach drug discovery, enabling them to leverage complex data sets and make informed decisions.
Machine Learning Frameworks and Libraries
Several popular machine learning frameworks and libraries are used in drug discovery, including TensorFlow and PyTorch. TensorFlow is an open-source software library developed by Google, widely used for large-scale machine learning tasks. It provides a high-level interface for building and training machine learning models, as well as low-level operations for building custom models.
PyTorch, on the other hand, is an open-source machine learning library developed by Facebook, known for its ease of use and rapid prototyping capabilities. Both TensorFlow and PyTorch have been adopted by the research community for their versatility and scalability.
Cloud Computing Platforms
Cloud computing platforms play a crucial role in drug discovery machine learning, providing efficient computation and storage needs. Cloud-based platforms such as Amazon Web Services (AWS), Microsoft Azure, and Google Cloud Platform (GCP) offer scalable computing power, data storage, and analytics services, making it possible to process large amounts of data efficiently.
By leveraging cloud computing platforms, researchers can access vast computing power, storage capacity, and advanced analytics tools, enabling them to analyze complex data sets, perform simulations, and make predictions with greater accuracy.
Open-Source Software and Tools
Several open-source software and tools are used in drug discovery machine learning, including RDKit and Biopython. RDKit is a software library for cheminformatics, used for molecular modeling, simulation, and analysis. It provides a range of tools for working with molecular structures, including molecular modeling, simulation, and analysis.
Biopython is a Python-based library for bioinformatics, used for molecular modeling, simulation, and analysis. It provides a range of tools for working with biological data, including molecular modeling, simulation, and analysis. Both RDKit and Biopython have been widely adopted by the research community for their versatility and scalability.
Other Tools and Technologies
In addition to machine learning frameworks, libraries, and software, several other tools and technologies are used in drug discovery machine learning, including:
*
- Deep learning libraries such as Keras and Caffe;
- Data pre-processing tools such as Pandas and NumPy;
- Data visualization tools such as Matplotlib and Seaborn;
- High-performance computing (HPC) clusters and GPU-accelerated computing;
- Cloud-based services such as AWS SageMaker and Google Cloud AI Platform.
These tools and technologies have revolutionized the way researchers approach drug discovery, enabling them to leverage complex data sets, make informed decisions, and develop new treatments with greater accuracy and speed.
Challenges and Opportunities in Drug Discovery Machine Learning
The rapid advancement of machine learning algorithms and large-scale data generation in drug discovery has led to numerous breakthroughs in this field. However, several challenges and opportunities continue to hinder or propel the progress of machine learning-driven drug discovery research. One of the significant challenges in this area is the complexity of molecular biology data and the lack of explainability in machine learning models.
Model Interpretability and Explainability Challenges
Machine learning models in drug discovery often employ complex algorithms that are difficult to interpret and explain. Molecular biology data, including protein structures, gene expression profiles, and patient outcomes, can be highly multidimensional and noisy. This data complexity poses significant challenges to developing models that can accurately predict drug efficacy, safety, and efficacy.
The lack of interpretability in machine learning models can lead to several issues, including:
- Lack of understanding of the underlying mechanisms driving drug efficacy and safety
- Difficulty in reproducing results in new datasets or different study populations
- Inability to identify potential biases in data and models
To address these challenges, researchers are developing new machine learning algorithms that prioritize interpretability and transparency, such as saliency maps, feature importance, and model agnostic explanations.
Diverse Datasets and Bias Mitigation
Another significant challenge in drug discovery machine learning is the need for diverse and representative datasets to avoid biases in models. Biased models can lead to poor generalizability and may not perform well in diverse populations or scenarios.
The lack of diverse datasets in drug discovery machine learning can be attributed to several factors, including:
- Limited access to clinical datasets and real-world patient data
- Biased representation of patients in clinical trials and studies
- Lack of standardization in data collection and reporting
To mitigate these biases, researchers are working on developing new data curation and collection strategies that prioritize diversity and inclusivity. This includes the use of synthetic data augmentation, transfer learning, and ensemble methods to improve model performance and generalizability.
Emerging Trends and Future Directions
Despite the challenges, machine learning-driven drug discovery research is rapidly advancing, with several emerging trends and future directions. Some of these include:
- Increased use of transfer learning and domain adaptation to leverage knowledge from other domains and data sources
- Development of hybrid models that combine the strengths of different machine learning algorithms (e.g., neural networks and decision trees)
- Integration of machine learning with other computational tools, such as molecular modeling, virtual screening, and systems biology
These emerging trends and future directions will likely lead to significant breakthroughs in drug discovery machine learning, enabling the development of more effective and personalized treatments for complex diseases.
As the complexity of molecular biology data continues to grow, machine learning algorithms will need to adapt and evolve to meet the demands of this rapidly changing field.
Future of Machine Learning in Drug Discovery
In the near future, machine learning is expected to revolutionize the field of drug discovery by facilitating the development of more effective treatments for various diseases. The integration of machine learning algorithms and techniques with large datasets and computational power will enable researchers to identify novel drug candidates, predict their efficacy, and streamline the drug development process.
Machine learning will play a pivotal role in drug discovery by leveraging vast amounts of data from various sources, such as genomics, proteomics, and clinical trials. By analyzing these data, researchers can gain insights into the underlying mechanisms of diseases and identify potential therapeutic targets. This will enable the design of more effective drugs that are tailored to individual patients’ needs.
Furthermore, machine learning can help reduce the time and cost associated with drug development by identifying potential failures early in the process. By analyzing large datasets and patterns, researchers can predict which compounds are more likely to succeed or fail, allowing for more targeted and efficient resource allocation.
Hypothetical Scenario: Breakthrough Discoveries in Disease Treatment
Let’s consider a hypothetical scenario where machine learning-driven drug discovery has led to breakthrough discoveries in disease treatment. In this scenario, researchers have developed a machine learning model that can analyze large datasets from various sources, including genomic data, clinical trials, and patient outcomes. Using this model, researchers have identified a novel compound that has shown incredible promise in treating a range of diseases, including cancer, Alzheimer’s, and diabetes.
This compound, which we’ll call “ML-001,” has been shown to have a high efficacy rate and minimal side effects. As a result, it has gained significant attention from the medical community, and several clinical trials have been initiated to test its safety and efficacy in human patients.
Collaborations between Experts from Computer Science, Biology, Pharmacology, and Medicine
The development of ML-001 required close collaboration between experts from various fields, including computer science, biology, pharmacology, and medicine. Computer scientists developed the machine learning model that analyzed the large datasets, while biologists and pharmacologists provided insights into the underlying mechanisms of disease and the potential therapeutic targets.
Pharmacologists played a critical role in designing the compound and optimizing its structure for maximum efficacy. Meanwhile, clinicians provided expertise on the clinical trials and helped to translate the findings into practical applications.
The collaboration was facilitated by the use of shared infrastructure and platforms that enabled the seamless exchange of data and ideas between researchers. This collaborative approach has been instrumental in enabling the development of more effective treatments and has set a new standard for interdisciplinary research in the field of drug discovery.
Advancements and Potential Applications of Machine Learning in Drug Discovery
Here are some of the key advancements and potential applications of machine learning in drug discovery:
| Method | Application | Example | Benefits |
|---|---|---|---|
| Deep Learning | Identification of novel therapeutic targets | Analysis of genomic data to identify potential targets for cancer treatment | Improved understanding of disease mechanisms and identification of new therapeutic opportunities |
| Reinforcement Learning | Optimization of compound structures for maximum efficacy | Use of machine learning algorithms to optimize the structure of ML-001 | Improved efficacy and reduced side effects |
| Natural Language Processing | Analysis of clinical trial data and literature to identify potential therapeutic opportunities | Use of NLP to analyze clinical trial data and identify patterns related to disease mechanisms | Improved understanding of disease mechanisms and identification of new therapeutic opportunities |
| Transfer Learning | Identification of potential therapeutic targets in related diseases | Use of transfer learning to identify potential targets for Alzheimer’s disease based on knowledge of Parkinson’s disease | Improved understanding of disease mechanisms and identification of new therapeutic opportunities |
In drug discovery, machine learning models are trained on high-dimensional data, which requires careful procedures to ensure accurate and reliable results. The training and validation processes involve feeding the model with labeled data, evaluating its performance, and refining it until it achieves satisfactory results. Here, we discuss the procedures for training and validating machine learning models on high-dimensional data, best practices for integrating multiple machine learning techniques, and step-by-step procedures for visualizing and analyzing the outputs of machine learning models in drug discovery applications.
Training and Validating Machine Learning Models, Drug discovery machine learning
The process of training machine learning models involves feeding the model with labeled data, which is a collection of input data and their corresponding output labels. The model learns patterns and relationships within the data to make predictions on new, unseen data. The quality of the labeled data has a direct impact on the performance of the model, and high-quality data is essential for training accurate models.
- Data Preprocessing: The first step in training a machine learning model is data preprocessing, which involves cleaning the data, handling missing values, and normalizing the features. This step is crucial in ensuring that the data is in a suitable format for the model to learn from.
- Model Selection: The choice of machine learning model depends on the type of problem being solved. For example, regression models are suitable for continuous targets, while classification models are suitable for categorical targets.
- Model Training: The trained model is evaluated on a validation dataset to assess its performance and identify areas for improvement.
The validation process involves splitting the data into training and validation sets, training the model on the training set, and evaluating its performance on the validation set.
Visualizing and Analyzing Machine Learning Model Outputs
Once the machine learning model has been trained and validated, it is essential to visualize and analyze its outputs to understand its behavior and limitations. The outputs can be visualized using techniques such as heatmaps, scatter plots, and confusion matrices.
| Method | Application | Example | Benefits |
|---|---|---|---|
| Heatmaps | Feature importance | A heatmap can be used to represent the importance of features in a machine learning model. For example, a heatmap can show the correlation between features and the target variable. | Heatmaps provide a visual representation of the data, making it easier to identify patterns and relationships. |
| Scatter plots | Data distribution | A scatter plot can be used to represent the distribution of the data. For example, a scatter plot can show the relationship between two continuous variables. | Scatter plots provide a visual representation of the data, making it easier to identify patterns and relationships. |
| Confusion matrices | Model performance | A confusion matrix can be used to evaluate the performance of a classification model. For example, a confusion matrix can show the true positives, false positives, true negatives, and false negatives. | Confusion matrices provide a summary of the model’s performance, making it easier to identify areas for improvement. |
“Data visualization is a very important aspect of machine learning. It helps to identify patterns and relationships that may not be apparent from the raw data.”
Integrating Multiple Machine Learning Techniques
Integrating multiple machine learning techniques can increase the performance of a machine learning model. For example, combining a linear regression model with a decision tree model can improve the accuracy of predictions.
- Ensemble Methods: Ensemble methods involve combining the predictions of multiple models to improve the overall performance. For example, a random forest model combines the predictions of multiple decision tree models.
- Stacking: Stacking involves combining the predictions of multiple models using a meta-model. For example, a linear regression model is used to combine the predictions of multiple decision tree models.
“Combining multiple machine learning techniques can improve the performance of a model by leveraging the strengths of each individual model.”
Final Summary
In conclusion, drug discovery machine learning holds immense promise for accelerating the development of new treatments and improving patient outcomes. While challenges remain, the integration of machine learning into the drug discovery process has the potential to revolutionize the pharmaceutical industry, making it more efficient, effective, and accessible.
Questions Often Asked
Q: What are the key challenges in traditional drug discovery methods?
The key challenges in traditional drug discovery methods include high costs, long development times, and a low success rate in identifying effective treatments.
Q: How can machine learning improve the drug discovery process?
Machine learning can improve the drug discovery process by analyzing large amounts of data, identifying patterns, and predicting the effectiveness of new treatments.
Q: What types of data are used in machine learning models for drug discovery?
The types of data used in machine learning models for drug discovery include genomic data, proteomic data, and chemical descriptors.